Use Case — Peer-Reviewed
AI-Powered Target Identification & Deconvolution
In a peer-reviewed study, Plex's AI platform identified novel drug targets from phenotypic screening data — predictions that were then validated in biochemical assays and a mouse model of liver disease, reducing hepatic steatosis by 68%.
Phenotypic Screening Coupled with AI-Driven Target Deconvolution Identifies α-Terthienyl as a Dual DPP-IV/HSD17β13 Modulator with Efficacy in a Mouse Model of MASLD
Read the paper →The Challenge: Target Deconvolution in Phenotypic Drug Discovery
Phenotypic screening identifies compounds that work in disease-relevant cell models — but it doesn't tell you why they work. Target deconvolution (identifying the molecular targets of a hit compound) traditionally requires months of pull-down assays, affinity chromatography, and genetic knockdown experiments. This bottleneck delays the transition from hit to lead and limits the ability to optimize compounds rationally.
How Plex Solved It
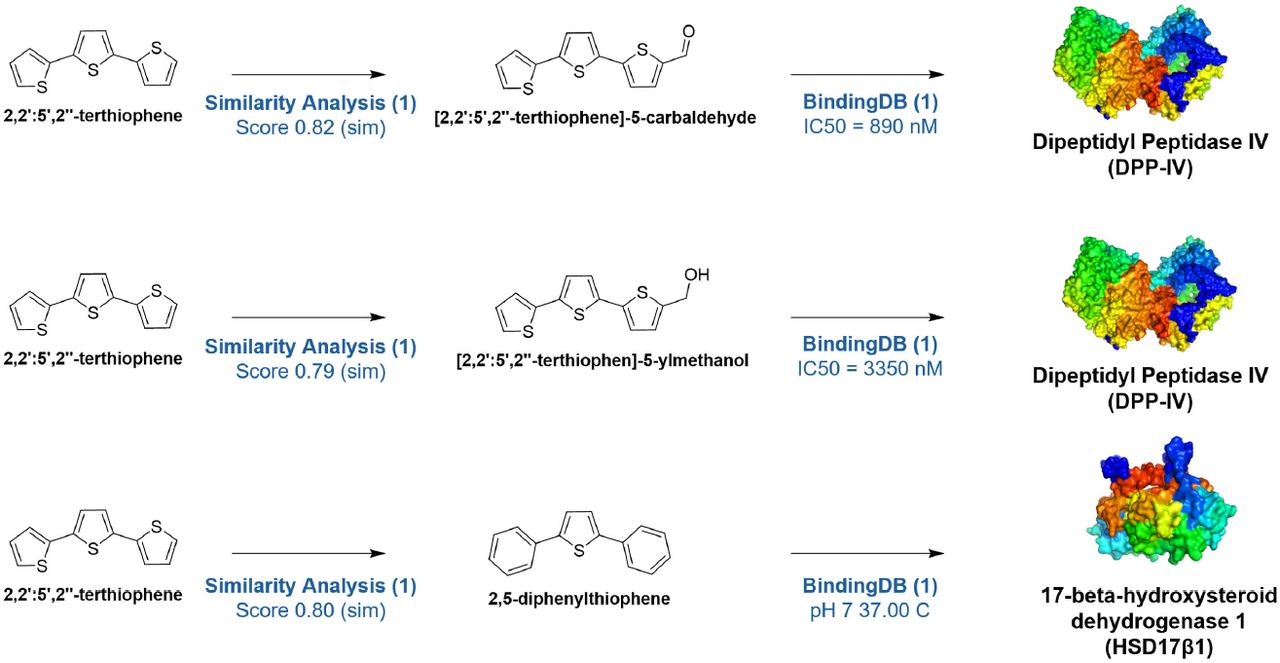
Researchers at the University of Michigan used Plex's AI platform for target deconvolution of their lead compound, α-terthienyl, identified from a high-content phenotypic screen for MASLD (fatty liver disease). Plex integrated the compound's chemical structure, quantitative phenotypic fingerprint, and biological knowledge graphs from 60+ public databases to generate focal graphs — knowledge subgraphs analyzed using centrality algorithms like PageRank — that ranked protein target hypotheses by probability.
What Plex Identified
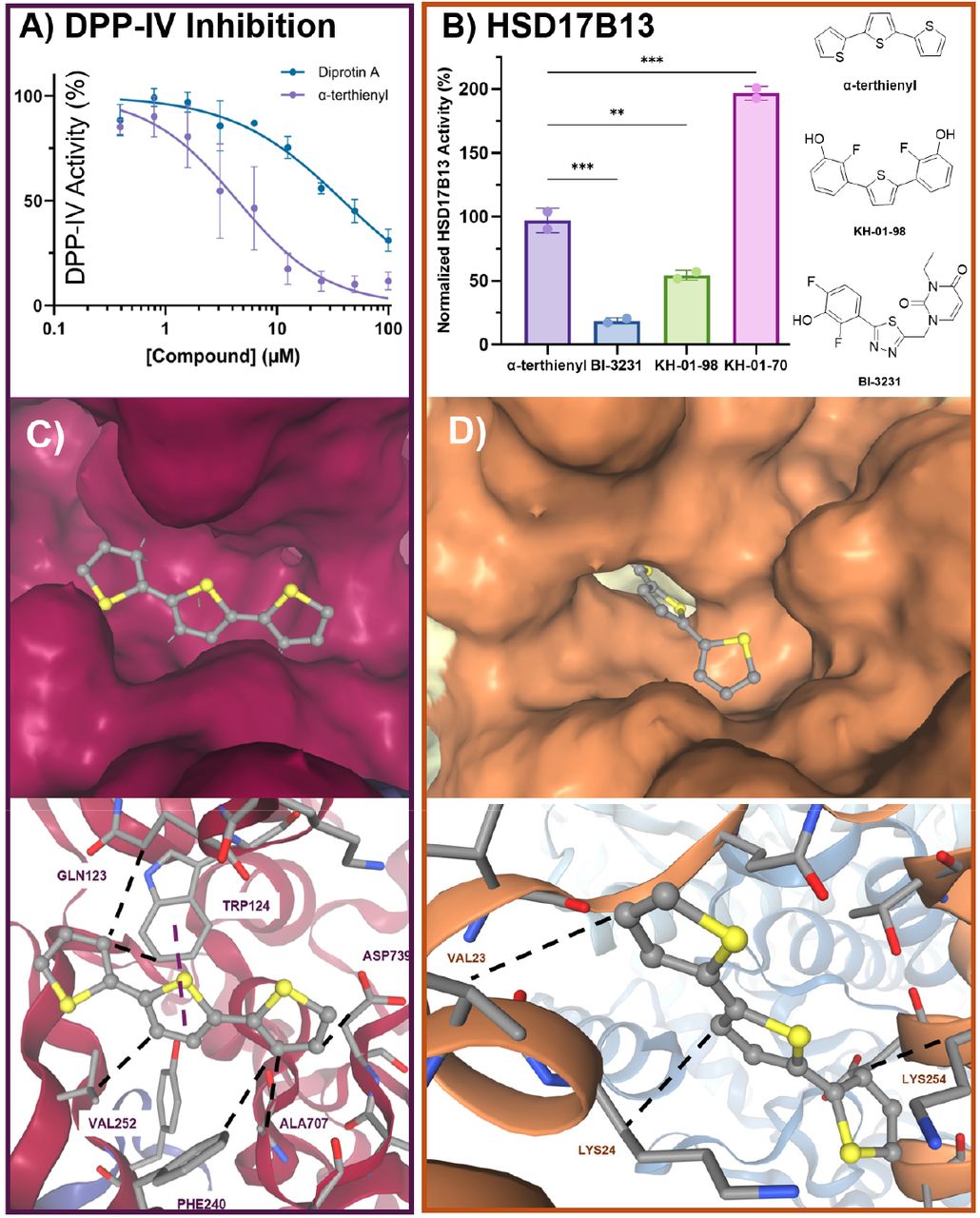
- DPP-IV (dipeptidyl peptidase 4) — a validated metabolic target, confirmed by biochemical assay with IC50 of 1.47 μM
- HSD17β13 (17-beta hydroxysteroid dehydrogenase 13) — a genetically validated MASLD target, confirmed by enzymatic inhibition assay
- Dual-target polypharmacology — Plex predicted the compound would act through both targets simultaneously, a mechanism validated in vivo
- 8 total target hypotheses ranked by confidence, with the top predictions confirmed experimentally
Validated Results
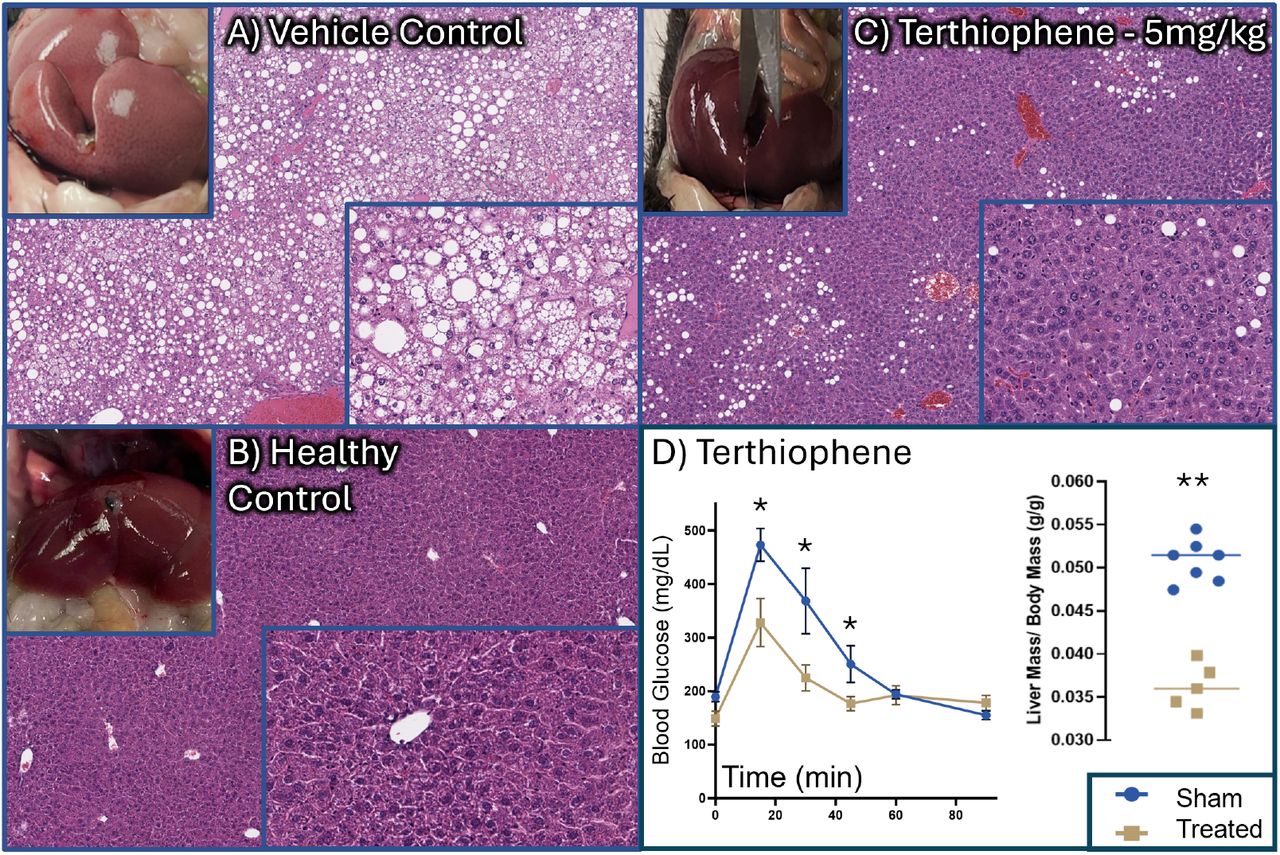
The Plex-predicted targets were validated across multiple experimental approaches: molecular docking confirmed binding affinity, biochemical assays confirmed enzymatic inhibition, and an in vivo mouse model demonstrated 68% reduction in hepatic steatosis, near-complete resolution of microvesicular steatosis, significantly improved glucose tolerance, and normalized liver biomarkers (ALT/AST). The compound showed an EC50 of 106 nM with no cytotoxicity up to 15 μM.
Key Capabilities Demonstrated
- AI-driven target deconvolution from phenotypic screening data — no prior target hypothesis required
- Multi-modal integration of chemical structures, phenotypic fingerprints, and biological knowledge graphs
- Focal Graph analysis using PageRank-style algorithms to rank target hypotheses by probability
- Rapid identification of polypharmacological mechanisms that would take months to discover experimentally
- Predictions validated by independent biochemical assays and in vivo disease models
See What Plex Can Identify in Your Therapeutic Area
Get a free report based on your science. Our AI will analyze your pipeline and surface novel target opportunities — the same platform validated in peer-reviewed research.
Get a Free Report Based on Your Science